SENTINEL: Difference between revisions
Veroandreo (talk | contribs) |
⚠️Griembauer (talk | contribs) (→Atmospheric correction with Atmospheric and Radiometric Correction of Satellite Imagery (ARCSI): Update of arcsi installation guide) |
||
| (34 intermediate revisions by 6 users not shown) | |||
| Line 36: | Line 36: | ||
* [https://github.com/ibamacsr/sentinelsat/ sentinelsat]: Python utilities to access the API of Copernicus Sentinels Scientific Data Hub (useful for all Sentinel satellite imagery) | * [https://github.com/ibamacsr/sentinelsat/ sentinelsat]: Python utilities to access the API of Copernicus Sentinels Scientific Data Hub (useful for all Sentinel satellite imagery) | ||
* [https://github.com/olivierhagolle/Sentinel-download/ Sentinel-download]: Automated download of Sentinel-2 L1C data from ESA (through wget) | * [https://github.com/olivierhagolle/Sentinel-download/ Sentinel-download]: Automated download of Sentinel-2 L1C data from ESA (through wget) | ||
'''Dedicated GRASS GIS Addons "i.sentinel.*":''' | |||
GRASS GIS has a dedicated set of {{AddonCmd|i.sentinel}} Addons, '''{{AddonCmd|i.sentinel.download}}''' and '''{{AddonCmd|i.sentinel.import}}''', to download and import Sentinel data into GRASS GIS database, respectively. This is probably the most convenient and straight-forward way if working with GRASS. Note that {{AddonCmd|i.sentinel.download}} is build on top of the ''sentinelsat'' library and it also requires ''python-pandas''. | |||
To install these addons, just run: | |||
<source lang="bash"> | |||
g.extension extension=i.sentinel | |||
</source> | |||
==Data pre-processing== | ==Data pre-processing== | ||
| Line 43: | Line 53: | ||
==Sentinel-2 in GRASS GIS== | ==Sentinel-2 in GRASS GIS== | ||
To exemplify the handling of Sentinel-2 data with GRASS GIS, we will use S2 L1C scene from North Carolina (NC) from November 2016. Here are the details: | |||
S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502 | |||
830f2a5a-9914-427c-b4e0-69c6c14c71d4 | |||
2016-11-10T15:05:02Z | |||
<!-- | |||
S2A_OPER_PRD_MSIL1C_PDMC_20161001T232624_R097_V20161001T160052_20161001T160426 | |||
44b7fb4c-d2d7-4739-9b8e-3639659505db | |||
2016-10-01T16:00:52.000Z | |||
--> | |||
In the zipped folder delivered by ESA, the data itself is hidden under: | |||
S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE/GRANULE/S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_N02.04/IMG_DATA/ | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B01.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B02.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B03.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B04.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B05.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B06.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B07.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B08.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B09.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B10.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B11.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B12.jp2 | |||
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B8A.jp2 | |||
<!-- explain naming --> | |||
===Importing footprints=== | |||
ESA has published a KML file containing the footprints (global coverage), see https://sentinel.esa.int/web/sentinel/missions/sentinel-2/data-products | |||
This file can be imported into a LongLat-WGS84 location: | |||
<source lang="bash"> | |||
# download KML | |||
wget https://sentinel.esa.int/documents/247904/1955685/S2A_OPER_GIP_TILPAR_MPC__20151209T095117_V20150622T000000_21000101T000000_B00.kml | |||
# create GRASS GIS location (or re-use an existing one) | |||
grass78 -c epsg:4326 ~/grassdata/longlat_wgs84 | |||
# now we are in the PERMANENT mapset | |||
# import KML without generating topology (-c flag) | |||
v.in.ogr S2A_OPER_GIP_TILPAR_MPC__20151209T095117_V20150622T000000_21000101T000000_B00.kml out=sentinel2_tileindex_global -c | |||
# done. | |||
</source> | |||
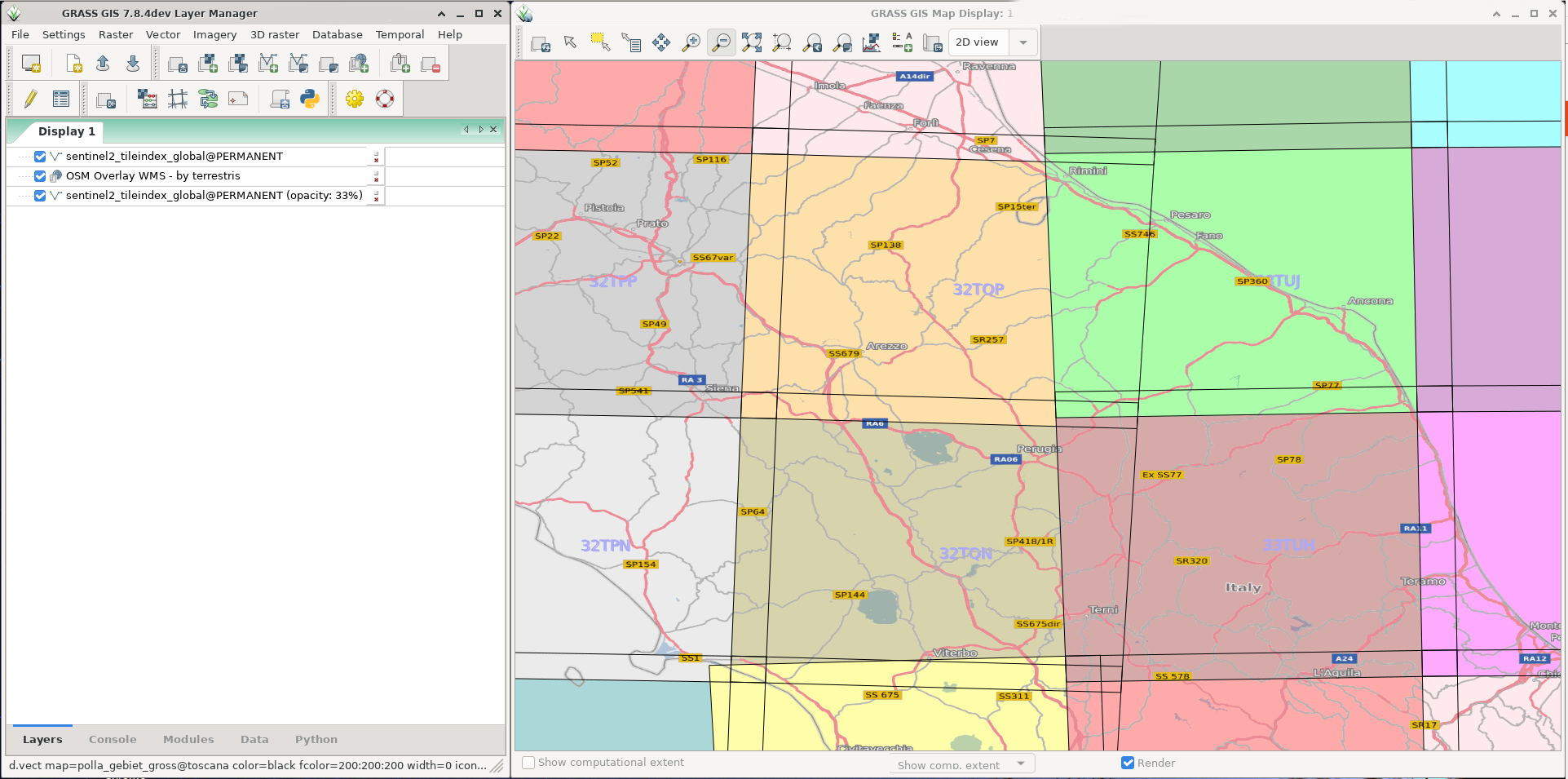
Next you can visualize the tile polygons and use the "Name" attribute column to show the tile labels (here, with OpenStreetMap WMS overlay from http://ows.terrestris.de/osm/service?): | |||
<center> | |||
[[File:Sentinel2 tiles.png|400px|Subset of Sentinel-2 tiles]] | |||
</center> | |||
===Importing data=== | |||
To import Sentinel-2 data into GRASS GIS, the simplest is to use the {{AddonCmd|i.sentinel.import}} addon. This nice module allows to list all files in a folder and import all the bands at once (with re-projection on the fly if needed), only import some of them by a pattern or just link the data through {{cmd|r.external}}. Here, are some examples on how to use i.sentinel.import: | |||
<source lang="bash"> | |||
# list all sentinel bands in a folder | |||
i.sentinel.import -p input=S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE memory=1000 | |||
# import all sentinel bands within the folder | |||
i.sentinel.import input=S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE memory=1000 | |||
# import only band 4 and 8 | |||
i.sentinel.import input=S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE memory=1000 pattern='B0[4|8]' | |||
# link files through r.external (-l flag) | |||
i.sentinel.import -l imput=S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE | |||
# import all bands with re-projection on the fly | |||
i.sentinel.import -r imput=S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE | |||
</source> | |||
Sentinel-2 data can be also easily imported into GRASS GIS by means of {{cmd|r.in.gdal}} (in a UTM location) or using {{cmd|r.import}} that provides re-projection on the fly. In this example, we will use {{cmd|r.import}} to re-project and import into the NC sample location delivered with GRASS GIS. | |||
<source lang="bash"> | |||
# list all jp2 files and import them | |||
for map in `ls *.jp2` ; do | |||
outmap=`basename $map .jp2` | |||
r.import input=$map output=$outmap | |||
done | |||
</source> | |||
If you prefer shorter map names, then it is possible to use {{cmd|g.rename}} or the addon {{AddonCmd|g.rename.many}}. The other option is editing the file names before importing with sed for example. Let us keep the S2A, date and band number parts only: | |||
<source lang="bash"> | |||
for map in `ls *.jp2` ; do | |||
outmap1=`basename $map .jp2` | |||
outmap2=`echo $outmap1 | sed s/OPER_MSI_L1C_TL_MTI__//g | sed s/T210803_A007244_T17SPV//g` | |||
r.import input=$map output=$outmap2 | |||
done | |||
</source> | |||
The output resolution estimated by {{cmd|r.import}} is slightly different than the nominal resolution of Sentinel-2 bands. | |||
<source lang="bash"> | |||
r.info S2A_20161110_B08 -g | grep res | |||
nsres=9.80178836938829 | |||
ewres=9.80224757766693 | |||
</source> | |||
We need to set the proper resolution at import by means of the '''resolution''' and '''resolution_value''' options in {{cmd|r.import}}. | |||
<source lang="bash"> | |||
for i in 02 03 04 08 ; do | |||
g.message message="importing band ${i} with R 10m" | |||
r.import S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B${i}.jp2 out=S2A_20161110_B${i} resolution=value resolution_value=10 extent=input | |||
done | |||
for i in 05 06 07 8A 11 12 ; do | |||
g.message message="importing band ${i} with R 20m" | |||
r.import S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B${i}.jp2 out=S2A_20161110_B${i} resolution=value resolution_value=20 extent=input | |||
done | |||
for i in 01 09 10 ; do | |||
g.message message="importing band ${i} with R 60m" | |||
r.import S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B${i}.jp2 out=S2A_20161110_B${i} resolution=value resolution_value=60 extent=input | |||
done | |||
</source> | |||
Hint concerning the black border around the scene data: you can use the "footprint" vector map which can be generated during the {{AddonCmd|i.sentinel.download}} step along with {{cmd|r.mask}} to suppress the black border (no-data area of a Sentinel-2 scene). | |||
In L2A products, there are artifacts along edges of swath paths, cell values are too low, caused by a bug in sen2cor. | |||
All cells that are bordering a NULL cell must be set to NULL. This step might need to be repeated twice. | |||
Because neighboring Sentinel tiles are overlapping each other, file sizes can be reduced by patching together neighboring tiles with | |||
the same datatake sensing start time. | |||
Before the actual re-projection, the computational region must be set properly. In the original projection, | |||
four 10 meter cells fall exactly into one 20 meter cell, and nine 20 meter cells fall exactly into one 60 meter cell. | |||
This can be achieved by first setting the region to the input extents (obtained with {{cmd|r.proj}} -g), then | |||
aligning the region to a resolution of 60 meter, after that setting the region's resolution to the actual resolution of | |||
the input band, this time without alignment. Alternatively, a reference grid can be created and the region for the current | |||
input can then be aligned to that reference grid. | |||
Now the data can be re-projected, preferably with method=bicubic_f or method=lanczos_f to avoid the sawtooth patterns | |||
of linear features which can appear with method=nearest (default). | |||
===Atmospheric correction with {{cmd|i.atcorr}}=== | ===Atmospheric correction with {{cmd|i.atcorr}}=== | ||
To exemplify the atmospheric correction of Sentinel-2 data, we download the L1C scene <code>S2A_OPER_PRD_MSIL1C_PDMC_20161029T092602_R054_V20161028T155402_20161028T155402</code> that overlaps completely with the ''elevation'' map in North Carolina location. This scene was imported with region cropping (see {{cmd|r.import}}) into PERMANENT mapset. The computational region was set to the extent of the ''elevation'' map. The atmospheric correction, as implemented by {{cmd|i.atcorr}}, is applied separately to each band of the scene and needs certain parameters to be set. | |||
In the first step we create a file containing the 6S parameters for a particular scene and band. To create a 6S file, we need to obtain the following information: | |||
* geometrical conditions, | |||
* moth, day, decimal hours in GMT, decimal longitude and latitude of measurement, | |||
* atmospheric model, | |||
* aerosol model, | |||
* visibility or aerosol optical depth, | |||
* mean target elevation above sea level, | |||
* sensor height and, | |||
* sensor band. | |||
Some of those details are encompassed in the metadata of the scene/bands and others are codified in different tables provided in the {{cmd|i.atcorr}} manual page. Let's go one by one: | |||
====Geometrical conditions==== | |||
For Sentinel-2A, the geometrical conditions take the value 25 and for Sentinel-2B, the geometrical conditions value is 26 (See table A in i.atcorr manual page). | |||
====Day, time, longitude and latitude of measurement==== | |||
Day and time of the measurement are hidden in the file name (i.e., the second datum in the file name with format YYYYMMDDTHHMMSS), and are also noted in the metadata file, which is included in the downloaded scene (file with .xml extension). Our sample scene was taken on October 28th (20161028) at 15:54:02 (155402). Note that the time has to be specified in decimal hours in Greenwich Mean Time (GMT). Luckily, the time in the scene name is in GMT and we can convert it to decimal hours as follows: 15 + 54/60 + 2/3600 = 15.901. | |||
Longitude and latitude refer to the centre of the computational region (which can be smaller than the scene), and must be in WGS84 decimal coordinates. To obtain the coordinates of the center, we can run: | |||
<source lang="bash"> | |||
g.region -bg | |||
</source> | |||
The longitude and latitude of the center are stored are <code>ll_clon=-78.691</code> and <code>ll_clat=35.749</code>. | |||
====Atmospheric model==== | |||
We can choose between various atmospheric models as defined at the beginning of this manual. For North Carolina, we can choose ''2 - midlatitude summer''. | |||
====Aerosol model==== | |||
We can also choose between various aerosol models as defined at the beginning of this manual. For North Carolina, we can choose ''1 - continental model''. | |||
====Visibility or Aerosol Optical Depth==== | |||
For Sentinel-2 scenes, the visibility is not measured, and therefore we have to estimate the aerosol optical depth instead, e.g. from [https://aeronet.gsfc.nasa.gov AERONET]. With a bit of luck, you can find a station nearby your location which measured the Aerosol Optical Depth at 500 nm at the same time as the scene was taken. In our case, on 28th October 2016, the ''EPA-Res_Triangle_Pk'' station measured AOD = 0.07 (approximately). | |||
====Mean target elevation above sea level==== | |||
Mean target elevation above sea level refers to the mean elevation of the computational region. You can estimate it from the digital elevation model, e.g. by running: | |||
<source lang="bash"> | |||
r.univar -g elevation | |||
</source> | |||
In our case, mean=110. In the 6S file it will be displayed in [-km], i.e., -0.110. | |||
====Sensor height==== | |||
Since the sensor is on board a satellite, the sensor height will be set to -1000. | |||
====Sensor band==== | |||
The overview of satellite bands can be found in table F in i.atcorr manual page. For Sentinel-2A, the band numbers span from 166 to 178, and for Sentinel-2B, from 179 to 191. | |||
Finally, here is what the 6S file would look like for Band 02 of our scene. In order to use it in the i.atcorr module, we can save it in a text file, for example <code>params_B02.txt</code>. | |||
25 | |||
10 28 15.901 -78.691 35.749 | |||
2 | |||
1 | |||
0 | |||
0.07 | |||
-0.110 | |||
-1000 | |||
167 | |||
Now, we compute the atmospheric correction for the selected band B02 of our Sentinel 2 scene. We have to specify the following parameters: | |||
* input = raster band to be processed, | |||
* parameters = path to 6S file created in the previous step (we could also enter the values directly), | |||
* output = name for the output corrected raster band, | |||
* range = from 1 to the QUANTIFICATION_VALUE stored in the metadata file. It is 10000 for both Sentinel-2A and Sentinel-2B. | |||
* rescale = the output range of values for the corrected bands. This is up to the user to choose, for example: 0-255, 0-1, 1-10000. | |||
If the data is available, the following parameters can be specified as well: | |||
* elevation = raster of digital elevation model, | |||
* visibility = raster of visibility model. | |||
Sentinel-2 Level 1C data represent TOA (top-of-atmosphere) reflectance values. By default, {{cmd|i.atcorr}} assumes that the input is TOA radiance, so the -r flag has to be set when running the command to indicate that the input is already in TOA reflectance. Finally, this is how the command would look like to apply atmospheric correction to band B02: | |||
<source lang="bash"> | |||
i.atcorr -r input=B02 parameters=params_B02.txt output=B02.atcorr range=1,10000 rescale=0,255 elevation=elevation | |||
</source> | |||
To apply atmospheric correction to the remaining bands, only the last line in the 6S parameters file (i.e., the sensor band) needs to be changed. The other parameters will remain the same. | |||
===Atmospheric correction with Atmospheric and Radiometric Correction of Satellite Imagery (ARCSI)=== | |||
The Atmospheric and Radiometric Correction of Satellite Imagery (ARCSI, https://github.com/remotesensinginfo/arcsi) software was developed at Aberystwyth University, under contract from the Norwegian Space Centre and allows automatic correction to Top Of Atmosphere (TOA) or Surface Reflectance (SREF). The conversion to Surface Reflectance uses the 6S code, accessed through the [http://py6s.rtwilson.com/ Py6S] Python interface. It was originally developed for Landsat data but it also supports Sentinel 2 satellites. | |||
Aerosol Optical Thickness (AOT) is an important parameter in atmospheric correction models such as 6S. In some regions of the world it has been found that a constant can be used. However, for most regions a constant is not viable as the atmosphere is very variable. Incorrect parametrization for AOT can lead to large errors in atmospheric correction. ARCSI provides a method of deriving AOT from an image by using a dark object subtraction (DOS) to estimate surface reflectance in the blue channel. Taking this estimated reflectance as input, 6S is then numerically inverted to identify an AOT value which provides a surface reflectance value as close to that estimated from DOS as possible. | |||
Instructions to install the software can be found at: https://arcsi.remotesensing.info/, but you can also have it through a docker image from: https://github.com/mundialis/docker-arcsi (for a usage example, see below). | |||
''Important'': The resulting atmospherically corrected satellite data are stored in '''KEA format''' (extended HDF) and can be converted with <tt>gdal_translate</tt> but ONLY if GDAL was compiled with http://www.kealib.org/ support or the kealib GDAL-plugin is installed! | |||
{| class="wikitable sortable" | |||
! ARCSI Product !! Definition | |||
|- | |||
| CLEARSKY || Identify large continuous areas without cloud, providing an image mask | |||
|- | |||
| CLOUDS || Perform a classification to mask cloud and cloud shadow | |||
|- | |||
| DDVAOT || Dark Dense Vegetation Aerosol Optical Thickness – used for atmospheric correction | |||
|- | |||
| DEM || Extract and reproject the inputted DEM for the image | |||
|- | |||
| DOS || Perform a dark object subtraction to retrieve an estimate of surface reflectance | |||
|- | |||
| DOSAOT || Estimate the Aerosol Optical Thickness (AOT) for the scene | |||
|- | |||
| DOSAOTSGL || Estimate a single value of Aerosol Optical Thickness (AOT) for the scene | |||
|- | |||
| FOOTPRINT || Extract the image data footprint as a shapefile | |||
|- | |||
| METADATA || Create a metadata file with key information from the input image and information calculated during the analysis | |||
|- | |||
| RAD || At sensor radiance image | |||
|- | |||
| SATURATE || Mask of saturated pixels | |||
|- | |||
| SHARP || Sharpen the 20 m image bands to 10 m using a local correlation filter | |||
|- | |||
| SREF || Using the 6S atmospheric model calculate the surface reflectance pixel value for the scene | |||
|- | |||
| STDSREF || Calculate standardised reflectance, normalising for the sun angle and topography (i.e., topographic correction) | |||
|- | |||
| THERMAL || Calculate thermal product (not for Sentinel-2) | |||
|- | |||
| TOA || At sensor reflectance (top of atmosphere; TOA) | |||
|- | |||
| TOPOSHADOW || Produce a mask for topographic shadow using the DEM | |||
|} | |||
''Sources:'' | |||
* "Developing a standards and automated production for Sentinel-2 Analysis ready Data - SD1707" ([http://sciencesearch.defra.gov.uk/Default.aspx?Menu=Menu&Module=More&Location=None&Completed=0&ProjectID=19836 ref]) | |||
* https://groups.google.com/g/rsgislib-support/c/CQ_S4BRuE3Y | |||
==== ARCSI: usage examples with DOS1 and DOSAOTSGL corrections ==== | |||
Here are some examples on how to use arcsi for Sentinel data. The first one only applies simple DOS1 correction and the second one demonstrates the application of the more advanced correction "DOSAOTSGL" which masks for clouds, cloud shadows and topographic shadows. The latter also derives AOT automatically. | |||
<source lang="bash"> | |||
# define name of Sentinel-2 scene - Note: omit .SAFE | |||
S2IMG=S2A_MSIL1C_20170327T105021_N0204_R051_T31UFT_20170327T105021 | |||
DEM=srtm_30m_myregion.tif | |||
OUTDIR=arcsi_output_AOT_inv | |||
TMPDIR=~/tmp/arcsi | |||
cd ${S2IMG}.SAFE/ | |||
mkdir ${TMPDIR} | |||
# simple DOS1 correction example | |||
arcsi.py --sensor sen2 -i MTD_MSIL1C.xml -o ${OUTDIR} \ | |||
--tmpath ${TMPDIR} -f KEA --stats -p RAD DOSAOTSGL SREF \ | |||
--aeroimg ${CONDA_PREFIX}/share/arcsi/WorldAerosolParams.kea \ | |||
--atmosimg ${CONDA_PREFIX}/share/arcsi/WorldAtmosphereParams.kea \ | |||
--dem ${DEM} --minaot 0.05 --maxaot 0.6 --simpledos | |||
# DOSAOTSGL correction example | |||
arcsi.py -s sen2 --stats --format KEA \ | |||
-p CLOUDS DOSAOTSGL STDSREF SATURATE TOPOSHADOW FOOTPRINT METADATA SHARP \ | |||
-o ${OUTDIR} --dem ${DEM} --tmpath ${TMPDIR} \ | |||
--k clouds.kea meta.json sat.kea toposhad.kea valid.kea stdsref.kea \ | |||
-i ${S2IMG}.SAFE/MTD_MSIL1C.xml | |||
</source> | |||
Where: | |||
* <code>-s</code> or <code>--sensor sen2</code> specifies the sensor, in the examples Sentinel-2 | |||
* <code>-i MTD_MSIL1C.xml</code> is the header file of the input image | |||
* <code>-o ${OUTDIR}</code> specifies the output directory where all output files will be saved to | |||
* <code>--tmpath ${TMPDIR}</code> is a directory where temporary files can be put during the processing; they will be deleted afterwards | |||
* <code>-f KEA</code> specifies that the output image file format should be KEA; any GDAL supported format can be used | |||
* <code>--stats</code> specifies that the output images should be populated with statistics and pyramids; makes display much faster but it is only available for KEA and HFA output formats | |||
* <code>-p</code> or <code>--prodlist RAD TOA SREF</code> specifies the output products to be generated (RAD: radiance, TOA: top-of-atmosphere, SREF: surface reflectance) | |||
* <code>--aeroimg ../WorldAerosolParams.kea</code> is an image file which is sampled to identify the aerosol model to be used for the input image (this can be downloaded from ARCSI downloads) | |||
* <code>--atmosimg ../WorldAtmosphereParams.kea</code> is an image file which is sampled to identify the atmosphere model to be used for the input image (this can be downloaded from ARCSI downloads) | |||
* <code>--dem ${DEM}</code> is an elevation model covering the scene. The higher the resolution of the DEM available the better | |||
* <code>--minaot 0.05</code> is the lower limit of the AOT values tested when estimating the AOT value to be used to correct the scene | |||
* <code>--maxaot 0.6</code> is the upper limit of the AOT values tested when estimating the AOT value to be used to correct the scene | |||
* <code>--simpledos</code> specifies that a simple single valued DOS method should be used to provide an estimate of the Blue SREF used in the inversion | |||
Further Sentinel-2 examples (taken from [https://github.com/mundialis/docker-arcsi mundialis github]): | |||
<source lang="bash"> | |||
# SENTINEL CLOUDS MASKING (ONLY) | |||
arcsi.py --sensor sen2 -i ${S2IMG}.SAFE/MTD_MSIL1C.xml -o ${S2IMG}.SAFE/Clouds \ | |||
--tmpath ${TMPDIR} -f KEA --stats -p CLOUDS | |||
# SENTINEL CLOUDS MASKING AND ATMCOR with 6S, with lookup of atmosphere profile | |||
arcsi.py --sensor sen2 -i ${S2IMG}.SAFE/MTD_MSIL1C.xml -o ${S2IMG}.SAFE/OutputsAOTInvCL \ | |||
--tmpath ${TMPDIR} -f KEA --stats -p CLOUDS RAD DOSAOTSGL SREF \ | |||
--aeroimg Documents/arcsi-1.4.2/data/WorldAerosolParams.kea \ | |||
--atmosimg Documents/arcsi-1.4.2/data/WorldAtmosphereParams.kea \ | |||
--dem ${S2IMG}.SAFE/dem_VR_all --minaot 0.05 --maxaot 0.6 --simpledos | |||
# SENTINEL CLOUDS MASKING AND ATMCOR with 6S but with fixed AOT (already known) | |||
arcsi.py --sensor sen2 -i ${S2IMG}.SAFE/MTD_MSIL1C.xml -o ${S2IMG}.SAFE/OutputsAOTInvCL \ | |||
--tmpath ${TMPDIR} -f KEA --stats -p CLOUDS SREF \ | |||
--aeroimg Documents/arcsi-1.4.2/data/WorldAerosolParams.kea \ | |||
--atmosimg Documents/arcsi-1.4.2/data/WorldAtmosphereParams.kea \ | |||
--dem ${S2IMG}.SAFE/dem_VR_all --aot 0.3 | |||
# SENTINEL, OLD NAME style from 2016; simple DOS example | |||
arcsi.py --sensor sen2 -i ${S2IMG}.SAFE/S2A_OPER_MTD_SAFL1C_PDMC_20170119T125545_R097_V20161120T160552_20161120T160552.xml \ | |||
-o ${S2IMG}.SAFE/OutputsAOTInv --tmpath ${TMPDIR} -f KEA --stats -p RAD DOSAOTSGL SREF \ | |||
--aeroimg Documents/arcsi-1.4.2/data/WorldAerosolParams.kea \ | |||
--atmosimg Documents/arcsi-1.4.2/data/WorldAtmosphereParams.kea \ | |||
--dem ${S2IMG}.SAFE/srtm_21_05_utm17 --minaot 0.05 --maxaot 0.6 --simpledos | |||
</source> | |||
All resulting files are stored in KEA format. | |||
==== ARCSI: usage example using docker ==== | |||
Using ARCSI based on docker image (download from https://hub.docker.com/r/mundialis/arcsi/) | |||
<source lang="bash"> | |||
# install ARCSI via docker | |||
docker pull mundialis/arcsi | |||
# define name of Sentinel-2 scene - note: omit: .SAFE | |||
S2IMG=S2A_MSIL1C_20170327T105021_N0204_R051_T31UFT_20170327T105021 | |||
DEM=srtm_30m_myregion.tif | |||
MY_S2_PATH=/path/to/S2_data/ | |||
MY_DEM_PATH=/path/to/DEM_data/ | |||
## ESA XML name: | |||
XML=MTD_MSIL1C.xml | |||
## USGS (modified XML name!): | |||
# XML=$(echo $S2IMG | sed 's+PRD_MSIL1C+MTD_SAFL1C+g') | |||
# run ARCSI (we use volume mapping to make S2 and DEM visible inside the docker container) | |||
# produce CLOUD and Surface Reflectance results, use DOSAOTSGL for AOT estimation | |||
docker run -it --rm -v ${MY_S2_PATH}:/data -v ${MY_DEM_PATH}:/dem mundialis/arcsi \ | |||
arcsi.py --sensor sen2 -i /data/${S2IMG}.SAFE/$XML.xml -o /data/${S2IMG}.SAFE/output \ | |||
--tmpath /tmp -f KEA --stats -p CLOUDS DOSAOTSGL SREF \ | |||
--aeroimg /opt/conda/share/arcsi/WorldAerosolParams.kea \ | |||
--atmosimg /opt/conda/share/arcsi/WorldAtmosphereParams.kea \ | |||
--dem /dem/${DEM} --demnodata 0 --minaot 0.05 --maxaot 0.6 | |||
</source> | |||
All resulting files are stored in KEA format. | |||
<!-- | |||
===Apply cloud mask=== | |||
Example here | Example here | ||
--> | |||
===Atmospheric correction with Maja=== | |||
MAJA (for Maccs-Atcor Joint Algorithm), is an atmospheric correction and cloud screening software. | |||
Currently, Maja allows the following processing steps: | |||
* Atmospheric correction | |||
* Cloud detection | |||
* Estimation of the Aerosol-Optical-Depth (AOT) | |||
* Correction of environmental- and slope-effects | |||
Maja is based on a multi-temporal method - allowing to refine the outputs using the previous inputs. | |||
* Source code and installation instructions: https://github.com/CNES/MAJA | |||
===Estimate indices=== | |||
Given that Sentinel-2 MSI comes with 3 red edge bands, it allows to estimate several other vegetation indices, aside from the most classic ones. There is a quite exhaustive list at the [http://www.sentinel-hub.com/eotaxonomy/indices?page=1 Sentinel Hub] website. Here, only some of them: | |||
=== | * [https://www.sentinel-hub.com/eoproducts/ndvi-normalized-difference-vegetation-index Normalized Difference Vegetation Index]: NDVI = (B08 - B04) / (B08 + B04) | ||
* [http://www.sentinel-hub.com/eoproducts/evi-enhanced-vegetation-index-0 Enhanced Vegetation Index]: EVI = 2.5*(B08 - B04) / (B08 + 6*B04 - 7.5*B02 + 1) | |||
* [https://www.sentinel-hub.com/eoproducts/sr-simple-ratio-index Simple Ratio Index]: SR = B8 / B4 | |||
* [http://www.sentinel-hub.com/eoproducts/arvi Atmospherically Resistant Vegetation Index]: ARVI = (B08 - (2*B04 - B02)) / (B08 + (2*B04 - B02)) | |||
* [https://www.sentinel-hub.com/eoproducts/savi-soil-adjusted-vegetation-index Soil Adjusted Vegetation Index]: SAVI = 1.5 * (B08 - B04) / (B08 + B04 + 0.5) | |||
* [https://www.sentinel-hub.com/eoproducts/red-edge-ndvi Red Edge NDVI]: NDVI-Re = (B08 - B06) / (B08 + B06) | |||
* [http://www.sentinel-hub.com/eoproducts/chl-red-edge-chlorophyll-red-edge Red Edge Chlorophyll Index]: Chl-Re = B05 / B08 | |||
* [http://www.sentinel-hub.com/eoproducts/ireci-inverted-red-edge-chlorophyll-index Inverted Red Edge Chlorophyll Index]: IReCI = (B07 - B04) * B06 / B05 | |||
* [http://www.sentinel-hub.com/eoproducts/gndvi-green-normalized-difference-vegetation-index Green Normalized Difference Vegetation Index]: GNDVI = (B08 - B03) / (B08 + B03) | |||
The following is an example of how to calculate NDVI with a simple mapcalc expression (assuming that bands were previously atmospherically corrected and they were rescaled to 1-10000): | |||
=== | <source lang="bash"> | ||
# define bands | |||
B4=S2A_20161110_B04 | |||
B8=S2A_20161110_B08 | |||
# NDVI | |||
r.mapcalc --o \ | |||
expression="NDVI = \ | |||
if(${B8} <= 10000 && ${B4} <= 10000, \ | |||
float(${B8} - ${B4}) / float(${B8} + ${B4}), null())" | |||
</source> | |||
Note that a long list of vegetation and water indices can be obtained with {{cmd|i.vi}} and {{AddonCmd|i.wi}}, respectively. | |||
===Classification=== | ===Classification=== | ||
As with other remote sensing data, we can use Sentinel data to perform a classification. Here, a quick example of an unsupervised classification: | |||
<source lang="bash"> | |||
# make imagery groups & subgroups: i.group | |||
# bands | |||
i.group group=sentinel subgroup=bands input=S2A_20161110_B02,S2A_20161110_B03,S2A_20161110_B04,S2A_20161110_B08 | |||
# indices | |||
i.group group=sentinel subgroup=indices input=NDVI,EVI | |||
# all | |||
i.group group=sentinel subgroup=all input=S2A_20161110_B02,S2A_20161110_B03,S2A_20161110_B04,S2A_20161110_B08,NDVI,EVI | |||
# spectral signatures for land cover types: i.cluster | |||
i.cluster group=sentinel subgroup=all signaturefile=s2_all_sig classes=7 iterations=50 min_size=100 reportfile=s2_all_7.txt --o | |||
i.cluster group=sentinel subgroup=bands signaturefile=s2_bands_sig classes=7 iterations=50 min_size=100 reportfile=s2_bands_7.txt --o | |||
i.cluster group=sentinel subgroup=indices signaturefile=s2_ind_sig classes=7 iterations=50 min_size=100 reportfile=s2_ind_7.txt --o | |||
# unsupervised classification: i.maxlik | |||
i.maxlik group=sentinel subgroup=all signaturefile=s2_all_sig output=s2_all_7 reject=s2_all_7_reject | |||
i.maxlik group=sentinel subgroup=bands sigfile=s2_bands_sig class=s2_bands_7 reject=s2_bands_7_reject | |||
i.maxlik group=sentinel subgroup=indices sigfile=s2_ind_sig class=s2_ind_7 reject=s2_ind_7_reject | |||
</source> | |||
==Further reading== | ==Further reading== | ||
IMAGICO.DE blog | |||
* [http://blog.imagico.de/sentinel-2-having-a-first-look-at-the-data-part-1/ First look at Sentinel data] | * Jena GRASS GIS Workshop: | ||
* [http://blog.imagico.de/who-is-moving-the-cheese-again/ Who is moving the cheese again?] | ** [https://training.gismentors.eu/grass-gis-workshop-jena/units/20.html Unit 20 - Sentinel downloader] | ||
** [https://training.gismentors.eu/grass-gis-workshop-jena/units/21.html Unit 21 - Sentinel spatio-temporal] | |||
** [https://training.gismentors.eu/grass-gis-workshop-jena/units/22.html Unit 22 - Spatio-temporal scripting] | |||
** [https://training.gismentors.eu/grass-gis-workshop-jena/units/23.html Unit 23 - Spatio-temporal parallelization] | |||
IMAGICO.DE blog: | |||
* [http://blog.imagico.de/sentinel-2-having-a-first-look-at-the-data-part-1/ First look at Sentinel data] (2016) | |||
* [http://blog.imagico.de/who-is-moving-the-cheese-again/ Who is moving the cheese again?] (2016) | |||
[[Category: Documentation]] | [[Category: Documentation]] | ||
Latest revision as of 13:50, 17 February 2021
Sentinel 2
Sentinel-2 is an Earth observation mission developed by ESA as part of the Copernicus Programme to perform terrestrial observations in support of services such as forest monitoring, land cover changes detection, and natural disaster management. It consists of two identical satellites, Sentinel-2A and Sentinel-2B. Details about the mission, user and technical guides as well as other documentation can be found at the ESA dedicated website [1] and a number of other sites.
Overview of Sentinel-2 mission capabilities
Sentinel-2 mission has the following features:
- Multi-spectral data with 13 bands in the visible, near infrared (NIR), and short wave infrared (SWIR).
- Systematic global coverage of land surfaces from 56° S to 84° N, coastal waters, and all of the Mediterranean Sea
- Revisiting every 5 days under the same viewing angles. At high latitudes, Sentinel-2 swath overlaps and some regions are observed twice or more every 5 days, but with different viewing angles.
- Spatial resolutions of 10 m (Bands 2, 3, 4 and 8 - Blue, Green, Red and NIR), 20 m (Bands 5, 6 and 7 - Red Edge bands, Band 8A - Narrow infrared and Bands 11 and 12 - SWIR) and 60 m (Bands 1 - Coastal aerosol, Band 9 - Water vapour and Band 10 - SWIR/Cirrus)
- 290 km field of view
- Free and open data policy
Sentinel-2 products
- Level-1B product provides radiometrically corrected imagery in Top-Of-Atmosphere (TOA) radiance values and in sensor geometry. Level-1B product is composed of an ensemble of granules that are 25 km across track (AC) by 23 km along track (AL). Additionally, this product includes the refined geometry which is used to generate the Level-1C product. Level-1B pixel coordinates refer to the centre of each pixel.
- Level-1C product is composed of 100 km² single tiles (ortho-images in UTM/WGS84 projection). The Level-1C product results from using a Digital Elevation Model (DEM) to project the image in cartographic coordinates. Per-pixel radiometric measurements are provided in TOA reflectances with all parameters to transform them into radiances. Level-1C products are resampled with a constant Ground Sampling Distance (GSD) of 10, 20 and 60 m depending on the native resolution of the different spectral bands. In Level-1C products, pixel coordinates refer to the upper left corner of the pixel. Before September 2016, Level-1C was delivered as multi-tile product.
- Level-2A product provides Bottom Of Atmosphere (BOA) reflectance images derived from the associated Level-1C products. Therefore, each Level-2A product is also a 100 km² tile in cartographic geometry (UTM/WGS84 projection). BOA reflectances in Level-2A products are scaled by a factor of 10000.
Data download
Sentinel-2 data might be obtained from the Copernicus Open Access Hub, either by using the online interactive interface and browsing to your area of interest or by means of the API Hub. There are several other sites from which it is possible to download Sentinel-2 imagery as well. Here, some of them:
- Sentinel-app
- Amazon S3
- Copernicus Data and Exploitation Platform – Deutschland (CODE-DE)
- mundialis
- RemotePixel
- Land Viewer
For those preferring or needing to download data via the API, there also exist several options:
- sentinelsat: Python utilities to access the API of Copernicus Sentinels Scientific Data Hub (useful for all Sentinel satellite imagery)
- Sentinel-download: Automated download of Sentinel-2 L1C data from ESA (through wget)
Dedicated GRASS GIS Addons "i.sentinel.*":
GRASS GIS has a dedicated set of i.sentinel Addons, i.sentinel.download and i.sentinel.import, to download and import Sentinel data into GRASS GIS database, respectively. This is probably the most convenient and straight-forward way if working with GRASS. Note that i.sentinel.download is build on top of the sentinelsat library and it also requires python-pandas.
To install these addons, just run:
g.extension extension=i.sentinel
Data pre-processing
As from May 2017, ESA started to provide Level 2A products for Europe and they will reprocess the archive during this year (2017). However, it is (still) possible to download Level-1C products and customize the atmospheric correction by using different algorithms (i.e.: 6S, 6SV, arcsi) and/or a different DEM (Sen2Cor uses STRM 90m by default).
Sentinel-2 in GRASS GIS
To exemplify the handling of Sentinel-2 data with GRASS GIS, we will use S2 L1C scene from North Carolina (NC) from November 2016. Here are the details:
S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502 830f2a5a-9914-427c-b4e0-69c6c14c71d4 2016-11-10T15:05:02Z
In the zipped folder delivered by ESA, the data itself is hidden under:
S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE/GRANULE/S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_N02.04/IMG_DATA/
S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B01.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B02.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B03.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B04.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B05.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B06.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B07.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B08.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B09.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B10.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B11.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B12.jp2 S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B8A.jp2
Importing footprints
ESA has published a KML file containing the footprints (global coverage), see https://sentinel.esa.int/web/sentinel/missions/sentinel-2/data-products
This file can be imported into a LongLat-WGS84 location:
# download KML
wget https://sentinel.esa.int/documents/247904/1955685/S2A_OPER_GIP_TILPAR_MPC__20151209T095117_V20150622T000000_21000101T000000_B00.kml
# create GRASS GIS location (or re-use an existing one)
grass78 -c epsg:4326 ~/grassdata/longlat_wgs84
# now we are in the PERMANENT mapset
# import KML without generating topology (-c flag)
v.in.ogr S2A_OPER_GIP_TILPAR_MPC__20151209T095117_V20150622T000000_21000101T000000_B00.kml out=sentinel2_tileindex_global -c
# done.
Next you can visualize the tile polygons and use the "Name" attribute column to show the tile labels (here, with OpenStreetMap WMS overlay from http://ows.terrestris.de/osm/service?):
Importing data
To import Sentinel-2 data into GRASS GIS, the simplest is to use the i.sentinel.import addon. This nice module allows to list all files in a folder and import all the bands at once (with re-projection on the fly if needed), only import some of them by a pattern or just link the data through r.external. Here, are some examples on how to use i.sentinel.import:
# list all sentinel bands in a folder
i.sentinel.import -p input=S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE memory=1000
# import all sentinel bands within the folder
i.sentinel.import input=S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE memory=1000
# import only band 4 and 8
i.sentinel.import input=S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE memory=1000 pattern='B0[4|8]'
# link files through r.external (-l flag)
i.sentinel.import -l imput=S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE
# import all bands with re-projection on the fly
i.sentinel.import -r imput=S2A_OPER_PRD_MSIL1C_PDMC_20161111T123605_R097_V20161110T160502_20161110T160502.SAFE
Sentinel-2 data can be also easily imported into GRASS GIS by means of r.in.gdal (in a UTM location) or using r.import that provides re-projection on the fly. In this example, we will use r.import to re-project and import into the NC sample location delivered with GRASS GIS.
# list all jp2 files and import them
for map in `ls *.jp2` ; do
outmap=`basename $map .jp2`
r.import input=$map output=$outmap
done
If you prefer shorter map names, then it is possible to use g.rename or the addon g.rename.many. The other option is editing the file names before importing with sed for example. Let us keep the S2A, date and band number parts only:
for map in `ls *.jp2` ; do
outmap1=`basename $map .jp2`
outmap2=`echo $outmap1 | sed s/OPER_MSI_L1C_TL_MTI__//g | sed s/T210803_A007244_T17SPV//g`
r.import input=$map output=$outmap2
done
The output resolution estimated by r.import is slightly different than the nominal resolution of Sentinel-2 bands.
r.info S2A_20161110_B08 -g | grep res
nsres=9.80178836938829
ewres=9.80224757766693
We need to set the proper resolution at import by means of the resolution and resolution_value options in r.import.
for i in 02 03 04 08 ; do
g.message message="importing band ${i} with R 10m"
r.import S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B${i}.jp2 out=S2A_20161110_B${i} resolution=value resolution_value=10 extent=input
done
for i in 05 06 07 8A 11 12 ; do
g.message message="importing band ${i} with R 20m"
r.import S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B${i}.jp2 out=S2A_20161110_B${i} resolution=value resolution_value=20 extent=input
done
for i in 01 09 10 ; do
g.message message="importing band ${i} with R 60m"
r.import S2A_OPER_MSI_L1C_TL_MTI__20161110T210803_A007244_T17SPV_B${i}.jp2 out=S2A_20161110_B${i} resolution=value resolution_value=60 extent=input
done
Hint concerning the black border around the scene data: you can use the "footprint" vector map which can be generated during the i.sentinel.download step along with r.mask to suppress the black border (no-data area of a Sentinel-2 scene).
In L2A products, there are artifacts along edges of swath paths, cell values are too low, caused by a bug in sen2cor. All cells that are bordering a NULL cell must be set to NULL. This step might need to be repeated twice.
Because neighboring Sentinel tiles are overlapping each other, file sizes can be reduced by patching together neighboring tiles with the same datatake sensing start time.
Before the actual re-projection, the computational region must be set properly. In the original projection, four 10 meter cells fall exactly into one 20 meter cell, and nine 20 meter cells fall exactly into one 60 meter cell. This can be achieved by first setting the region to the input extents (obtained with r.proj -g), then aligning the region to a resolution of 60 meter, after that setting the region's resolution to the actual resolution of the input band, this time without alignment. Alternatively, a reference grid can be created and the region for the current input can then be aligned to that reference grid.
Now the data can be re-projected, preferably with method=bicubic_f or method=lanczos_f to avoid the sawtooth patterns of linear features which can appear with method=nearest (default).
Atmospheric correction with i.atcorr
To exemplify the atmospheric correction of Sentinel-2 data, we download the L1C scene S2A_OPER_PRD_MSIL1C_PDMC_20161029T092602_R054_V20161028T155402_20161028T155402 that overlaps completely with the elevation map in North Carolina location. This scene was imported with region cropping (see r.import) into PERMANENT mapset. The computational region was set to the extent of the elevation map. The atmospheric correction, as implemented by i.atcorr, is applied separately to each band of the scene and needs certain parameters to be set.
In the first step we create a file containing the 6S parameters for a particular scene and band. To create a 6S file, we need to obtain the following information:
- geometrical conditions,
- moth, day, decimal hours in GMT, decimal longitude and latitude of measurement,
- atmospheric model,
- aerosol model,
- visibility or aerosol optical depth,
- mean target elevation above sea level,
- sensor height and,
- sensor band.
Some of those details are encompassed in the metadata of the scene/bands and others are codified in different tables provided in the i.atcorr manual page. Let's go one by one:
Geometrical conditions
For Sentinel-2A, the geometrical conditions take the value 25 and for Sentinel-2B, the geometrical conditions value is 26 (See table A in i.atcorr manual page).
Day, time, longitude and latitude of measurement
Day and time of the measurement are hidden in the file name (i.e., the second datum in the file name with format YYYYMMDDTHHMMSS), and are also noted in the metadata file, which is included in the downloaded scene (file with .xml extension). Our sample scene was taken on October 28th (20161028) at 15:54:02 (155402). Note that the time has to be specified in decimal hours in Greenwich Mean Time (GMT). Luckily, the time in the scene name is in GMT and we can convert it to decimal hours as follows: 15 + 54/60 + 2/3600 = 15.901. Longitude and latitude refer to the centre of the computational region (which can be smaller than the scene), and must be in WGS84 decimal coordinates. To obtain the coordinates of the center, we can run:
g.region -bg
The longitude and latitude of the center are stored are ll_clon=-78.691 and ll_clat=35.749.
Atmospheric model
We can choose between various atmospheric models as defined at the beginning of this manual. For North Carolina, we can choose 2 - midlatitude summer.
Aerosol model
We can also choose between various aerosol models as defined at the beginning of this manual. For North Carolina, we can choose 1 - continental model.
Visibility or Aerosol Optical Depth
For Sentinel-2 scenes, the visibility is not measured, and therefore we have to estimate the aerosol optical depth instead, e.g. from AERONET. With a bit of luck, you can find a station nearby your location which measured the Aerosol Optical Depth at 500 nm at the same time as the scene was taken. In our case, on 28th October 2016, the EPA-Res_Triangle_Pk station measured AOD = 0.07 (approximately).
Mean target elevation above sea level
Mean target elevation above sea level refers to the mean elevation of the computational region. You can estimate it from the digital elevation model, e.g. by running:
r.univar -g elevation
In our case, mean=110. In the 6S file it will be displayed in [-km], i.e., -0.110.
Sensor height
Since the sensor is on board a satellite, the sensor height will be set to -1000.
Sensor band
The overview of satellite bands can be found in table F in i.atcorr manual page. For Sentinel-2A, the band numbers span from 166 to 178, and for Sentinel-2B, from 179 to 191.
Finally, here is what the 6S file would look like for Band 02 of our scene. In order to use it in the i.atcorr module, we can save it in a text file, for example params_B02.txt.
25 10 28 15.901 -78.691 35.749 2 1 0 0.07 -0.110 -1000 167
Now, we compute the atmospheric correction for the selected band B02 of our Sentinel 2 scene. We have to specify the following parameters:
- input = raster band to be processed,
- parameters = path to 6S file created in the previous step (we could also enter the values directly),
- output = name for the output corrected raster band,
- range = from 1 to the QUANTIFICATION_VALUE stored in the metadata file. It is 10000 for both Sentinel-2A and Sentinel-2B.
- rescale = the output range of values for the corrected bands. This is up to the user to choose, for example: 0-255, 0-1, 1-10000.
If the data is available, the following parameters can be specified as well:
- elevation = raster of digital elevation model,
- visibility = raster of visibility model.
Sentinel-2 Level 1C data represent TOA (top-of-atmosphere) reflectance values. By default, i.atcorr assumes that the input is TOA radiance, so the -r flag has to be set when running the command to indicate that the input is already in TOA reflectance. Finally, this is how the command would look like to apply atmospheric correction to band B02:
i.atcorr -r input=B02 parameters=params_B02.txt output=B02.atcorr range=1,10000 rescale=0,255 elevation=elevation
To apply atmospheric correction to the remaining bands, only the last line in the 6S parameters file (i.e., the sensor band) needs to be changed. The other parameters will remain the same.
Atmospheric correction with Atmospheric and Radiometric Correction of Satellite Imagery (ARCSI)
The Atmospheric and Radiometric Correction of Satellite Imagery (ARCSI, https://github.com/remotesensinginfo/arcsi) software was developed at Aberystwyth University, under contract from the Norwegian Space Centre and allows automatic correction to Top Of Atmosphere (TOA) or Surface Reflectance (SREF). The conversion to Surface Reflectance uses the 6S code, accessed through the Py6S Python interface. It was originally developed for Landsat data but it also supports Sentinel 2 satellites.
Aerosol Optical Thickness (AOT) is an important parameter in atmospheric correction models such as 6S. In some regions of the world it has been found that a constant can be used. However, for most regions a constant is not viable as the atmosphere is very variable. Incorrect parametrization for AOT can lead to large errors in atmospheric correction. ARCSI provides a method of deriving AOT from an image by using a dark object subtraction (DOS) to estimate surface reflectance in the blue channel. Taking this estimated reflectance as input, 6S is then numerically inverted to identify an AOT value which provides a surface reflectance value as close to that estimated from DOS as possible.
Instructions to install the software can be found at: https://arcsi.remotesensing.info/, but you can also have it through a docker image from: https://github.com/mundialis/docker-arcsi (for a usage example, see below).
Important: The resulting atmospherically corrected satellite data are stored in KEA format (extended HDF) and can be converted with gdal_translate but ONLY if GDAL was compiled with http://www.kealib.org/ support or the kealib GDAL-plugin is installed!
| ARCSI Product | Definition |
|---|---|
| CLEARSKY | Identify large continuous areas without cloud, providing an image mask |
| CLOUDS | Perform a classification to mask cloud and cloud shadow |
| DDVAOT | Dark Dense Vegetation Aerosol Optical Thickness – used for atmospheric correction |
| DEM | Extract and reproject the inputted DEM for the image |
| DOS | Perform a dark object subtraction to retrieve an estimate of surface reflectance |
| DOSAOT | Estimate the Aerosol Optical Thickness (AOT) for the scene |
| DOSAOTSGL | Estimate a single value of Aerosol Optical Thickness (AOT) for the scene |
| FOOTPRINT | Extract the image data footprint as a shapefile |
| METADATA | Create a metadata file with key information from the input image and information calculated during the analysis |
| RAD | At sensor radiance image |
| SATURATE | Mask of saturated pixels |
| SHARP | Sharpen the 20 m image bands to 10 m using a local correlation filter |
| SREF | Using the 6S atmospheric model calculate the surface reflectance pixel value for the scene |
| STDSREF | Calculate standardised reflectance, normalising for the sun angle and topography (i.e., topographic correction) |
| THERMAL | Calculate thermal product (not for Sentinel-2) |
| TOA | At sensor reflectance (top of atmosphere; TOA) |
| TOPOSHADOW | Produce a mask for topographic shadow using the DEM |
Sources:
- "Developing a standards and automated production for Sentinel-2 Analysis ready Data - SD1707" (ref)
- https://groups.google.com/g/rsgislib-support/c/CQ_S4BRuE3Y
ARCSI: usage examples with DOS1 and DOSAOTSGL corrections
Here are some examples on how to use arcsi for Sentinel data. The first one only applies simple DOS1 correction and the second one demonstrates the application of the more advanced correction "DOSAOTSGL" which masks for clouds, cloud shadows and topographic shadows. The latter also derives AOT automatically.
# define name of Sentinel-2 scene - Note: omit .SAFE
S2IMG=S2A_MSIL1C_20170327T105021_N0204_R051_T31UFT_20170327T105021
DEM=srtm_30m_myregion.tif
OUTDIR=arcsi_output_AOT_inv
TMPDIR=~/tmp/arcsi
cd ${S2IMG}.SAFE/
mkdir ${TMPDIR}
# simple DOS1 correction example
arcsi.py --sensor sen2 -i MTD_MSIL1C.xml -o ${OUTDIR} \
--tmpath ${TMPDIR} -f KEA --stats -p RAD DOSAOTSGL SREF \
--aeroimg ${CONDA_PREFIX}/share/arcsi/WorldAerosolParams.kea \
--atmosimg ${CONDA_PREFIX}/share/arcsi/WorldAtmosphereParams.kea \
--dem ${DEM} --minaot 0.05 --maxaot 0.6 --simpledos
# DOSAOTSGL correction example
arcsi.py -s sen2 --stats --format KEA \
-p CLOUDS DOSAOTSGL STDSREF SATURATE TOPOSHADOW FOOTPRINT METADATA SHARP \
-o ${OUTDIR} --dem ${DEM} --tmpath ${TMPDIR} \
--k clouds.kea meta.json sat.kea toposhad.kea valid.kea stdsref.kea \
-i ${S2IMG}.SAFE/MTD_MSIL1C.xml
Where:
-sor--sensor sen2specifies the sensor, in the examples Sentinel-2-i MTD_MSIL1C.xmlis the header file of the input image-o ${OUTDIR}specifies the output directory where all output files will be saved to--tmpath ${TMPDIR}is a directory where temporary files can be put during the processing; they will be deleted afterwards-f KEAspecifies that the output image file format should be KEA; any GDAL supported format can be used--statsspecifies that the output images should be populated with statistics and pyramids; makes display much faster but it is only available for KEA and HFA output formats-por--prodlist RAD TOA SREFspecifies the output products to be generated (RAD: radiance, TOA: top-of-atmosphere, SREF: surface reflectance)--aeroimg ../WorldAerosolParams.keais an image file which is sampled to identify the aerosol model to be used for the input image (this can be downloaded from ARCSI downloads)--atmosimg ../WorldAtmosphereParams.keais an image file which is sampled to identify the atmosphere model to be used for the input image (this can be downloaded from ARCSI downloads)--dem ${DEM}is an elevation model covering the scene. The higher the resolution of the DEM available the better--minaot 0.05is the lower limit of the AOT values tested when estimating the AOT value to be used to correct the scene--maxaot 0.6is the upper limit of the AOT values tested when estimating the AOT value to be used to correct the scene--simpledosspecifies that a simple single valued DOS method should be used to provide an estimate of the Blue SREF used in the inversion
Further Sentinel-2 examples (taken from mundialis github):
# SENTINEL CLOUDS MASKING (ONLY)
arcsi.py --sensor sen2 -i ${S2IMG}.SAFE/MTD_MSIL1C.xml -o ${S2IMG}.SAFE/Clouds \
--tmpath ${TMPDIR} -f KEA --stats -p CLOUDS
# SENTINEL CLOUDS MASKING AND ATMCOR with 6S, with lookup of atmosphere profile
arcsi.py --sensor sen2 -i ${S2IMG}.SAFE/MTD_MSIL1C.xml -o ${S2IMG}.SAFE/OutputsAOTInvCL \
--tmpath ${TMPDIR} -f KEA --stats -p CLOUDS RAD DOSAOTSGL SREF \
--aeroimg Documents/arcsi-1.4.2/data/WorldAerosolParams.kea \
--atmosimg Documents/arcsi-1.4.2/data/WorldAtmosphereParams.kea \
--dem ${S2IMG}.SAFE/dem_VR_all --minaot 0.05 --maxaot 0.6 --simpledos
# SENTINEL CLOUDS MASKING AND ATMCOR with 6S but with fixed AOT (already known)
arcsi.py --sensor sen2 -i ${S2IMG}.SAFE/MTD_MSIL1C.xml -o ${S2IMG}.SAFE/OutputsAOTInvCL \
--tmpath ${TMPDIR} -f KEA --stats -p CLOUDS SREF \
--aeroimg Documents/arcsi-1.4.2/data/WorldAerosolParams.kea \
--atmosimg Documents/arcsi-1.4.2/data/WorldAtmosphereParams.kea \
--dem ${S2IMG}.SAFE/dem_VR_all --aot 0.3
# SENTINEL, OLD NAME style from 2016; simple DOS example
arcsi.py --sensor sen2 -i ${S2IMG}.SAFE/S2A_OPER_MTD_SAFL1C_PDMC_20170119T125545_R097_V20161120T160552_20161120T160552.xml \
-o ${S2IMG}.SAFE/OutputsAOTInv --tmpath ${TMPDIR} -f KEA --stats -p RAD DOSAOTSGL SREF \
--aeroimg Documents/arcsi-1.4.2/data/WorldAerosolParams.kea \
--atmosimg Documents/arcsi-1.4.2/data/WorldAtmosphereParams.kea \
--dem ${S2IMG}.SAFE/srtm_21_05_utm17 --minaot 0.05 --maxaot 0.6 --simpledos
All resulting files are stored in KEA format.
ARCSI: usage example using docker
Using ARCSI based on docker image (download from https://hub.docker.com/r/mundialis/arcsi/)
# install ARCSI via docker
docker pull mundialis/arcsi
# define name of Sentinel-2 scene - note: omit: .SAFE
S2IMG=S2A_MSIL1C_20170327T105021_N0204_R051_T31UFT_20170327T105021
DEM=srtm_30m_myregion.tif
MY_S2_PATH=/path/to/S2_data/
MY_DEM_PATH=/path/to/DEM_data/
## ESA XML name:
XML=MTD_MSIL1C.xml
## USGS (modified XML name!):
# XML=$(echo $S2IMG | sed 's+PRD_MSIL1C+MTD_SAFL1C+g')
# run ARCSI (we use volume mapping to make S2 and DEM visible inside the docker container)
# produce CLOUD and Surface Reflectance results, use DOSAOTSGL for AOT estimation
docker run -it --rm -v ${MY_S2_PATH}:/data -v ${MY_DEM_PATH}:/dem mundialis/arcsi \
arcsi.py --sensor sen2 -i /data/${S2IMG}.SAFE/$XML.xml -o /data/${S2IMG}.SAFE/output \
--tmpath /tmp -f KEA --stats -p CLOUDS DOSAOTSGL SREF \
--aeroimg /opt/conda/share/arcsi/WorldAerosolParams.kea \
--atmosimg /opt/conda/share/arcsi/WorldAtmosphereParams.kea \
--dem /dem/${DEM} --demnodata 0 --minaot 0.05 --maxaot 0.6
All resulting files are stored in KEA format.
Atmospheric correction with Maja
MAJA (for Maccs-Atcor Joint Algorithm), is an atmospheric correction and cloud screening software.
Currently, Maja allows the following processing steps:
- Atmospheric correction
- Cloud detection
- Estimation of the Aerosol-Optical-Depth (AOT)
- Correction of environmental- and slope-effects
Maja is based on a multi-temporal method - allowing to refine the outputs using the previous inputs.
- Source code and installation instructions: https://github.com/CNES/MAJA
Estimate indices
Given that Sentinel-2 MSI comes with 3 red edge bands, it allows to estimate several other vegetation indices, aside from the most classic ones. There is a quite exhaustive list at the Sentinel Hub website. Here, only some of them:
- Normalized Difference Vegetation Index: NDVI = (B08 - B04) / (B08 + B04)
- Enhanced Vegetation Index: EVI = 2.5*(B08 - B04) / (B08 + 6*B04 - 7.5*B02 + 1)
- Simple Ratio Index: SR = B8 / B4
- Atmospherically Resistant Vegetation Index: ARVI = (B08 - (2*B04 - B02)) / (B08 + (2*B04 - B02))
- Soil Adjusted Vegetation Index: SAVI = 1.5 * (B08 - B04) / (B08 + B04 + 0.5)
- Red Edge NDVI: NDVI-Re = (B08 - B06) / (B08 + B06)
- Red Edge Chlorophyll Index: Chl-Re = B05 / B08
- Inverted Red Edge Chlorophyll Index: IReCI = (B07 - B04) * B06 / B05
- Green Normalized Difference Vegetation Index: GNDVI = (B08 - B03) / (B08 + B03)
The following is an example of how to calculate NDVI with a simple mapcalc expression (assuming that bands were previously atmospherically corrected and they were rescaled to 1-10000):
# define bands
B4=S2A_20161110_B04
B8=S2A_20161110_B08
# NDVI
r.mapcalc --o \
expression="NDVI = \
if(${B8} <= 10000 && ${B4} <= 10000, \
float(${B8} - ${B4}) / float(${B8} + ${B4}), null())"
Note that a long list of vegetation and water indices can be obtained with i.vi and i.wi, respectively.
Classification
As with other remote sensing data, we can use Sentinel data to perform a classification. Here, a quick example of an unsupervised classification:
# make imagery groups & subgroups: i.group
# bands
i.group group=sentinel subgroup=bands input=S2A_20161110_B02,S2A_20161110_B03,S2A_20161110_B04,S2A_20161110_B08
# indices
i.group group=sentinel subgroup=indices input=NDVI,EVI
# all
i.group group=sentinel subgroup=all input=S2A_20161110_B02,S2A_20161110_B03,S2A_20161110_B04,S2A_20161110_B08,NDVI,EVI
# spectral signatures for land cover types: i.cluster
i.cluster group=sentinel subgroup=all signaturefile=s2_all_sig classes=7 iterations=50 min_size=100 reportfile=s2_all_7.txt --o
i.cluster group=sentinel subgroup=bands signaturefile=s2_bands_sig classes=7 iterations=50 min_size=100 reportfile=s2_bands_7.txt --o
i.cluster group=sentinel subgroup=indices signaturefile=s2_ind_sig classes=7 iterations=50 min_size=100 reportfile=s2_ind_7.txt --o
# unsupervised classification: i.maxlik
i.maxlik group=sentinel subgroup=all signaturefile=s2_all_sig output=s2_all_7 reject=s2_all_7_reject
i.maxlik group=sentinel subgroup=bands sigfile=s2_bands_sig class=s2_bands_7 reject=s2_bands_7_reject
i.maxlik group=sentinel subgroup=indices sigfile=s2_ind_sig class=s2_ind_7 reject=s2_ind_7_reject
Further reading
- Jena GRASS GIS Workshop:
IMAGICO.DE blog:
- First look at Sentinel data (2016)
- Who is moving the cheese again? (2016)