R statistics/rgrass
Installation
Terminology
See "Overview" in R_statistics
R within GRASS
In this example, we will use R within a GRASS GIS session, i.e. start R (or RStudio) from the GRASS GIS terminal or command line interface.
Startup
- First start a GRASS GIS session. Then, at the GRASS command line start R (for an 'rstudio' session, see below)
- In this example we will use the North Carolina sample dataset.
Reset the region settings to the defaults
GRASS (nc_spm_08_grass7):~ > g.region -d
Launch R from the GRASS prompt
GRASS (nc_spm_08_grass7):~ > R
Load the rgrass library:
library(rgrass)Get the GRASS environment (mapset, region, map projection, etc.); you can display the metadata for your location by printing G:
gmeta()
gisdbase /home/mneteler/grassdata
location nc_spm_08_grass7
mapset user1
rows 620
columns 1630
north 320000
south 10000
west 120000
east 935000
nsres 500
ewres 500
projection:
PROJCRS["NAD83(HARN) / North Carolina",
BASEGEOGCRS["NAD83(HARN)",
DATUM["NAD83 (High Accuracy Reference Network)",
ELLIPSOID["GRS 1980",6378137,298.257222101,
LENGTHUNIT["metre",1]]],
PRIMEM["Greenwich",0,
ANGLEUNIT["degree",0.0174532925199433]],
ID["EPSG",4152]],
CONVERSION["SPCS83 North Carolina zone (meters)",
METHOD["Lambert Conic Conformal (2SP)",
ID["EPSG",9802]],
PARAMETER["Latitude of false origin",33.75,
ANGLEUNIT["degree",0.0174532925199433],
ID["EPSG",8821]],
PARAMETER["Longitude of false origin",-79,
ANGLEUNIT["degree",0.0174532925199433],
ID["EPSG",8822]],
PARAMETER["Latitude of 1st standard parallel",36.1666666666667,
ANGLEUNIT["degree",0.0174532925199433],
ID["EPSG",8823]],
PARAMETER["Latitude of 2nd standard parallel",34.3333333333333,
ANGLEUNIT["degree",0.0174532925199433],
ID["EPSG",8824]],
PARAMETER["Easting at false origin",609601.22,
LENGTHUNIT["metre",1],
ID["EPSG",8826]],
PARAMETER["Northing at false origin",0,
LENGTHUNIT["metre",1],
ID["EPSG",8827]]],
CS[Cartesian,2],
AXIS["easting (X)",east,
ORDER[1],
LENGTHUNIT["metre",1]],
AXIS["northing (Y)",north,
ORDER[2],
LENGTHUNIT["metre",1]],
USAGE[
SCOPE["Engineering survey, topographic mapping."],
AREA["United States (USA) - North Carolina - counties of Alamance; Alexander; Alleghany; Anson; Ashe; Avery; Beaufort; Bertie; Bladen; Brunswick; Buncombe; Burke; Cabarrus; Caldwell; Camden; Carteret; Caswell; Catawba; Chatham; Cherokee; Chowan; Clay; Cleveland; Columbus; Craven; Cumberland; Currituck; Dare; Davidson; Davie; Duplin; Durham; Edgecombe; Forsyth; Franklin; Gaston; Gates; Graham; Granville; Greene; Guilford; Halifax; Harnett; Haywood; Henderson; Hertford; Hoke; Hyde; Iredell; Jackson; Johnston; Jones; Lee; Lenoir; Lincoln; Macon; Madison; Martin; McDowell; Mecklenburg; Mitchell; Montgomery; Moore; Nash; New Hanover; Northampton; Onslow; Orange; Pamlico; Pasquotank; Pender; Perquimans; Person; Pitt; Polk; Randolph; Richmond; Robeson; Rockingham; Rowan; Rutherford; Sampson; Scotland; Stanly; Stokes; Surry; Swain; Transylvania; Tyrrell; Union; Vance; Wake; Warren; Washington; Watauga; Wayne; Wilkes; Wilson; Yadkin; Yancey."],
BBOX[33.83,-84.33,36.59,-75.38]],
ID["EPSG",3358]]Listing of existing maps
List available vector maps:
execGRASS("g.list", parameters = list(type = "vector"))List selected vector maps (wildcard):
execGRASS("g.list", parameters = list(type = "vector", pattern = "precip*"))
precip
precip_30ynormals
precip_30ynormals_3dSave selected vector maps into R vector:
my_vmaps <- execGRASS("g.list", parameters = list(type = "vector", pattern = "precip*"))
attributes(my_vmaps)
attributes(my_vmaps)$resOut
[1] "precip" "precip_30ynormals" "precip_30ynormals_3d"List available raster maps:
execGRASS("g.list", parameters = list(type = "raster"))List selected raster maps (wildcard):
execGRASS("g.list", parameters = list(type = "raster", pattern = "lsat7_2002*"))
lsat7_2002_10
lsat7_2002_20
lsat7_2002_30
lsat7_2002_40
lsat7_2002_50
lsat7_2002_61
lsat7_2002_62
lsat7_2002_70
lsat7_2002_80Reading data from GRASS
Example 1: raster maps
Manual page: read_RAST()
Read two GRASS raster maps, "geology_30m" and "elevation", from the North Carolina sample data location into the R current session as a new object "ncdata". This object is a SpatRaster layer from terra package:
ncdata <- read_RAST(c("geology_30m", "elevation"))We can verify the new R object:
ncdata
class : SpatRaster
dimensions : 450, 500, 2 (nrow, ncol, nlyr)
resolution : 30, 30 (x, y)
extent : 630000, 645000, 215000, 228500 (xmin, xmax, ymin, ymax)
coord. ref. : +proj=lcc +lat_0=33.75 +lon_0=-79 +lat_1=36.1666666666667 +lat_2=34.3333333333333 +x_0=609601.22 +y_0=0 +ellps=GRS80 +units=m +no_defs
sources : file1da12a752d65.grd
file1da12112d9ea4.grd
names : geology_30m, elevation
min values : 217, 55.92321
max values : 948, 156.25197Now, let's plot one of the raster layers within the SpatRaster object. We'll load terra library to have all functionality available:
library(terra)
plot(ncdata$elevation, main="North Carolina elevation")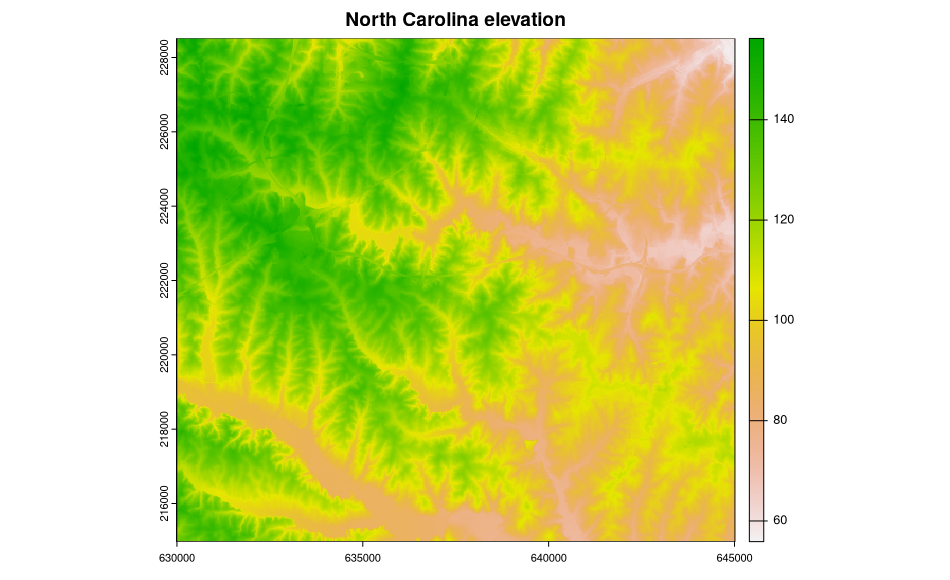
In addition, we can show what is going on inside the objects read into R:
summary(ncdata)
geology_30m elevation
Min. :217.0 Min. : 56.01
1st Qu.:217.0 1st Qu.: 94.77
Median :270.0 Median :108.86
Mean :311.2 Mean :110.36
3rd Qu.:270.0 3rd Qu.:126.80
Max. :948.0 Max. :156.07Example 2: single raster map
Here an example to transfer a single raster map from GRASS GIS to R:
library(rgrass)
library(terra)
execGRASS("g.region", raster = "elevation", flags = "p")
ncdata <- read_RAST("elevation")
summary(ncdata)
plot(ncdata)Example 3: vector maps
Here an example to transfer a single vector map from GRASS GIS to R. In this case, vector maps from GRASS will be converted into SpatVector objects from the terra package in R:
# needed prerequisite
library(rgrass)
library(terra)
execGRASS("v.info", map="schools_wake", layer="1")
vInfo("schools_wake")
myschools <- read_VECT("schools_wake")
print(summary(myschools))
plot(myschools)Example 4: Summarizing data
We can create a table of cell counts:
freq(ncdata$geology_30m)layer value count 1 1 217 80618 2 1 262 22076 3 1 270 76597 4 1 405 28190 5 1 583 2401 6 1 720 536 7 1 766 786 8 1 862 6858 9 1 910 4996 10 1 921 1392 11 1 945 1 12 1 946 452 13 1 948 97
And compare with the equivalent GRASS module:
execGRASS("r.stats", flags=c("c", "l"), parameters=list(input="geology_30m"), ignore.stderr=TRUE)217 CZfg 80618 262 CZlg 22076 270 CZig 76597 405 CZbg 28190 583 CZve 2401 720 CZam 536 766 CZg 786 862 CZam 6858 910 CZbg 4996 921 Km 1392 945 CZbg 1 946 CZam 452 948 CZam 97
Example 5: Querying maps
Sometimes you may want to query GRASS GIS maps from your R session. As an example, here the transfer of raster pixel values at the position of vector points:
# set the computational region to the raster map:
execGRASS("g.region", raster = "elev_state_500m", flags = "p")
# query raster map at vector points, transfer result into R
goutput <- execGRASS("r.what", map="elev_state_500m", points="geodetic_pts", separator=",", intern=TRUE)
str(goutput)
chr [1:29939] "571530.81289275,265739.968425953,,187.8082200648" ...
# parse it
con <- textConnection(goutput)
go1 <- read.csv(con, header=FALSE)
str(go1)
'data.frame': 29939 obs. of 4 variables:
$ V1: num 571531 571359 571976 572391 573011 ...
$ V2: num 265740 265987 267049 267513 269615 ...
$ V3: logi NA NA NA NA NA NA ...
$ V4: Factor w/ 22738 levels "-0.0048115728",..: 6859 6642 6749 6411 6356 6904 7506 7224 6908 7167 ...
summary(go1)
V1 V2 V3 V4
Min. :121862 Min. : 7991 Mode:logical 0 : 723
1st Qu.:462706 1st Qu.:162508 NA's:29939 * : 293
Median :610328 Median :204502 0.3048 : 47
Mean :588514 Mean :200038 0.6096 : 44
3rd Qu.:717610 3rd Qu.:247633 1.524 : 42
Max. :946743 Max. :327186 0.9144 : 23
(Other):28767Exporting data back to GRASS
Finally, a SpatialGridDataFrame object is written back to a GRASS raster map:
First prepare some data: (square root of elevation)
ncdata$sqdem <- sqrt(ncdata$elevation)Export data from R back into a GRASS raster map:
write_RAST(ncdata$sqdem, "sqdemNC", flags = c("o","overwrite"))Check that it imported properly:
execGRASS("r.info", parameters=list(map="sqdemNC"))+----------------------------------------------------------------------------+ | Map: sqdemNC Date: Fri Apr 21 17:14:05 2023 | | Mapset: PERMANENT Login of Creator: veroandreo | | Location: nc_spm_08_grass7 | | DataBase: /home/veroandreo/grassdata | | Title: | | Timestamp: none | |----------------------------------------------------------------------------| | | | Type of Map: raster Number of Categories: 0 | | Data Type: FCELL Semantic label: (none) | | Rows: 450 | | Columns: 500 | | Total Cells: 225000 | | Projection: Lambert Conformal Conic | | N: 228500 S: 215000 Res: 30 | | E: 645000 W: 630000 Res: 30 | | Range of data: min = 7.478182 max = 12.50008 | | | | Data Description: | | generated by r.in.gdal | | | | Comments: | | r.in.gdal --overwrite -o input="/home/veroandreo/tmp/grass8-veroandr\ | | eo-120898/RtmpPcRHkF/file1da12dbc5b5d.grd" output="sqdemNC" memory=3\ | | 00 offset=0 num_digits=0 | | | +----------------------------------------------------------------------------+
Using RStudio in a GRASS GIS session
If you are most used to run R through RStudio, but still want to have all GRASS data available for performing any analyses without loosing the possibility of still using GRASS command line, you can run:
GRASS> rstudio &
or, if you already are working on a certain RStudio project:
GRASS> rstudio /path/to/project/folder/ &
Then, you load rgrass library in your RStudio project
library(rgrass)and you are ready to go.
GRASS within R
GRASS within R
In this case, we start GRASS GIS within an R session, i.e., we use initGRASS() to start GRASS in a temporary location or in an existing GRASS GIS location.
The first argument in initGRASS() is gisBase=. This argument refers to the path where GRASS is installed. In the latest rgrass version, if the gisBase argument is missing, initGRASS() will do a minimal effort of guessing it. Firstly, if a GRASS_INSTALLATION environment variable is availabe, then its value will automatically be used for gisBase. If not, then the system command grass --config path</> will be tried to get the value of GISBASE that corresponds to the GRASS installation in the system PATH (if any).
If users, in any case want to pass the path to GRASS installation themselves, they can run:
# OSGeo4W shell, Linux, Mac OSX users:
grass --config path
It may report something like:
# OSGeo4W users:
C:\OSGeo4W64\apps\grass\grass
# Linux, Mac OSX users:
/usr/local/grass
If GRASS GIS was installed through OSGeo4W, users should first start the OSGeo4W Shell. Note that it may start in C: or a directory where the user does not have writing permission. Before calling R or RStudio, first change to a directory where you are sure you have writing permission.
## MS-Windows users (example for an OSGeo4W 64bit installation)
# change to a directory with writing permission
C:\>d:
D:\>cd temp
D:\temp>cd testR
# start R within this directory:
D:\temp\testR>R
# to start RStudio from OSGeo4W Shell, run: "C:/Program Files/RStudio/bin/rstudio.exe"
# load rgrass package
library(rgrass)
# initialisation of GRASS in the North Carolina sample dataset
initGRASS(gisBase = "C:/OSGeo4W64/apps/grass/grass",
gisDbase = "C:/Users/marissa/grassdata/",
location = "nc_spm_08_grass7",
mapset = "user1",
home=tempdir(),
override = TRUE)Linux and Mac OSX users just need to start R or RStudio the usual way, load the rgrass package and initialize GRASS as shown below:
library(rgrass)
# initialisation of GRASS in the North Carolina sample dataset
initGRASS(gisBase = "/usr/lib64/grass82",
gisDbase = "/home/veroandreo/grassdata/",
location = "nc_basic_spm_grass7",
mapset = "user1",
home=tempdir(),
override = TRUE)
Temporary location
If we don’t pass an existing location to initGRASS(), a temporary GRASS location will be created and set. By default this will be an XY location, i.e., no CRS defined. To define a CRS for the temporary location, we create a terra SpatRaster object that by default will be EPSG:4326.
x <- rast()
x
class : SpatRaster
dimensions : 180, 360, 1 (nrow, ncol, nlyr)
resolution : 1, 1 (x, y)
extent : -180, 180, -90, 90 (xmin, xmax, ymin, ymax)
coord. ref. : lon/lat WGS 84Then, we use that object to provide an EPSG to our temporary location via the SG= parameter and start GRASS:
initGRASS(home = tempdir(),
SG = x,
override = TRUE)After that, users can write raster and vector maps into GRASS with write_RAST() and write_VECT(), use GRASS GIS modules over those data with execGRASS() and export the results back to R with read_RAST() and read_VECT() as exemplified above.
Existing location
In the case of existing locations, users will provide the path to them as shown before and read data from GRASS database into R through read_RAST() and read_VECT(), do their analyses or modeling in R and potentially write results back into GRASS with write_RAST() and write_VECT().
R in GRASS in batch mode
Run the following script with
R CMD BATCH batch.R
library(rgrass)
# initialisation of GRASS in the North Carolina sample dataset (example with a compiled version)
initGRASS(gisBase = "/home/veroandreo/software/grass/dist.x86_64-pc-linux-gnu",
home = tempdir(),
gisDbase = "/home/veroandreo/grassdata/",
location = "nc_spm_08_grass7", mapset = "user1", SG="elevation",
override = TRUE)
# set region to default
system("g.region -dp")
# verify
gmeta()
# read data into R
ncdata <- read_RAST(c("geology_30m", "elevation"), cat=c(TRUE, FALSE))
# summary of geology map
summary(ncdata$geology_30m)
proc.time()The result is (shorted here):
cat batch.Rout
R version 4.0.2 (2020-06-22) -- "Taking Off Again"
Copyright (C) 2020 The R Foundation for Statistical Computing
Platform: x86_64-redhat-linux-gnu (64-bit)
...
> library(rgrass)
Loading required package: XML
GRASS GIS interface loaded with GRASS version: (GRASS not running)
> # initialisation and the use of north carolina dataset
> initGRASS(gisBase = "/home/veroandreo/software/grass/dist.x86_64-pc-linux-gnu", home = tempdir(),
+ gisDbase = "/home/veroandreo/grassdata/",
+ location = "nc_spm_08_grass7", mapset = "user1", SG="elevation",
+ override = TRUE)
gisdbase /home/veroandreo/grassdata/
location nc_spm_08_grass7
mapset user1
rows 13500
columns 15000
north 228500
south 215000
west 630000
east 645000
nsres 1
ewres 1
projection +proj=lcc +lat_1=36.16666666666666 +lat_2=34.33333333333334
+lat_0=33.75 +lon_0=-79 +x_0=609601.22 +y_0=0 +no_defs +a=6378137
+rf=298.257222101 +towgs84=0.000,0.000,0.000 +type=crs +to_meter=1
> system("g.region -dp")
projection: 99 (Lambert Conformal Conic)
zone: 0
datum: nad83
ellipsoid: a=6378137 es=0.006694380022900787
north: 320000
south: 10000
west: 120000
east: 935000
nsres: 500
ewres: 500
rows: 620
cols: 1630
cells: 1010600
> gmeta()
gisdbase /home/veroandreo/grassdata/
location nc_spm_08_grass7
mapset user1
rows 620
columns 1630
north 320000
south 10000
...
> ncdata <- read_RAST(c("geology_30m", "elevation"), cat=c(TRUE, FALSE))
...
> summary(ncdata$geology_30m)
CZfg_217 CZlg_262 CZig_270 CZbg_405 CZve_583 CZam_720 CZg_766 CZam_862
292 78 277 102 8 1 2 25
CZbg_910 Km_921 CZam_946 NA's
18 5 2 1009790
> proc.time()
user system elapsed
8.061 2.016 10.048Troubleshooting
Running out of disk space
Linux: A common issue is that /tmp/ is used for temporary files albeit being often a small partition. To change that to a larger directory, you may add to your $HOME/.bashrc the entry:
# change TMP directory of R (note: of course also another directory than suggested here is fine)
mkdir -p $HOME/tmp
export TMP=$HOME/tmp
The drawback of this approach is that on modern Linux systems the /tmp/ is a fast RAM disk (based on tempfs) while HOME directories are often on slower spinning disks (unless you have a SSD drive). At least you no longer run out of disk space easily.